ABRF 2026 — Pittsburgh, Pennsylvania
Multi-Site Benchmarking Studies For PRECYSE, A Novel DNA/RNA Single Molecule Imaging, Analysis and Sizing Platform For NGS Library QC
Introduction
Quality control (QC) remains a fundamental component of complex DNA sequencing workflows, ensuring the integrity of nucleic acid samples prior to downstream analysis, and validating intermediate constructs during library construction. Despite its importance, current QC technologies employed have not kept pace with the rapid evolution of sequencing platforms and applications, presenting a barrier to broader research and clinical implementation. In this study, we report findings from an extensive benchmarking evaluation of a novel biomolecular imaging and analysis system, PRECYSE, designed for rapid sizing and visualization of DNA fragments ranging from <100 to 500,000+ bp. Conducted across six prominent genomics core facilities in the United States, the study assessed over 200 DNA and RNA samples representing a broad spectrum of sample types and quality. These included short- and long-read sequencing libraries, gDNA derived from blood, saliva, and FFPE tissue, ultra-high molecular weight DNA for Oxford Nanopore sequencing, size-selected cfDNA, enzyme-bound libraries, and sheared genomic DNA.
Workflow
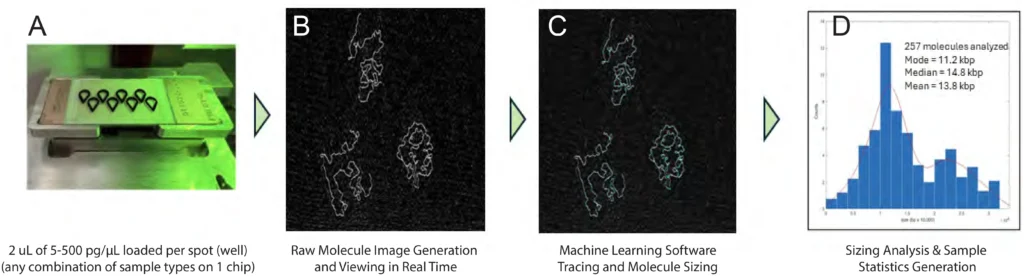

Figure 1. PRECYSE Workflow Overview
PRECYSE invokes proprietary scanning based on HSAFM (High Speed Atomic Force Microscopy) technology for direct imaging from 70 bp to 500+ kbp DNA
with single molecule resolution. Several plexes of the EZchip consumable are available up to 96 samples whereby each ‘spot’ requires 2 μL of DNA sample
ranging from 5 – 500 pg/μL (higher amount for larger DNA). Samples incubate for 5 min. followed by an automated wash, drying and heating step (not shown), a
17 min. total time from sample to scanning with no cold consumables.
(A) EZchip shown in PRECYSE instrument ready for scanning. Each sample spot is scanned sequentially, and data analyzed (DNA <20 kbp is a 2 min. scan to data generation time).
(B) Raw image output is viewed in real-time during scanning as partial ‘frames’. Shown here is a zoomed-in section of 1 frame showing 3 PacBio library molecules.
(C) Molecules are analyzed and sized by Molecular Explorer software in real-time during scanning. Molecule backbone tracing is shown in blue
(D) Molecules analyzed per sample are pooled together to generate a sizing histogram with sample statistics. Individual molecules can be interrogated and sizing ranges for statistical analysis can be modified.
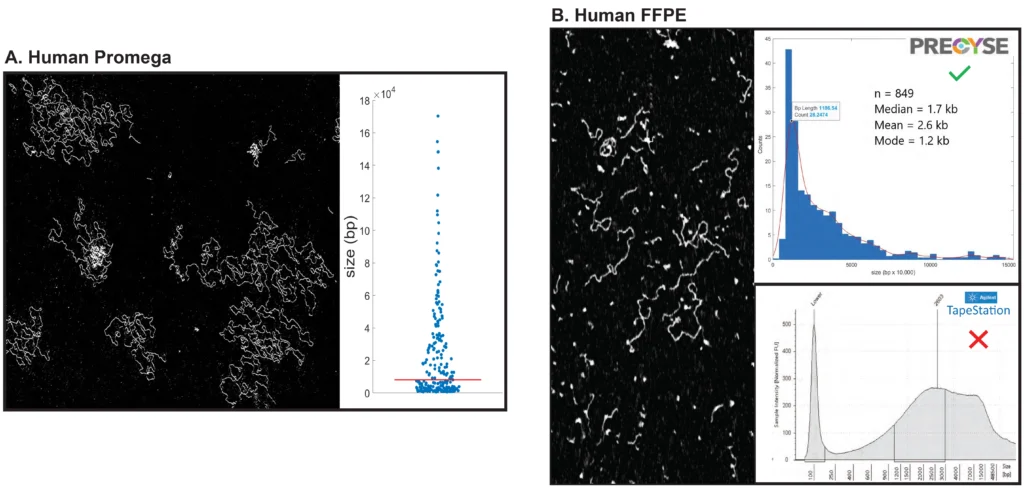
Figure 2. gDNA Analysis
(A) Stock female Promega gDNA scanned by PRECYSE at VCU. Partial raw image of one frame scan. Scatter plot sizing for 227 molecules shows a long distribution of larger molecules out to ~200 kb, indicative of intact high quality gDNA. Significant amount of ssDNA was observed.
(B) Partial raw image scan of Human gDNA extracted from FFPE and analyzed by The Jackson Laboratory. PRECYSE sizing analysis of 849 molecules generated a median size of 1.7 kb ranging from 212 to 19,784 bp, indicative of degraded/sheared gDNA. Extremely small DNA or particulates, and cross-linked DNA (intense thicker white molecules), was observed. The Agilent TapeStation electropherogram of same sample produced sizing from 250 to >48,500 bp (max instrument resolution). This larger incorrect sizing is presumably due to the highly crosslinked nature of FFPE gDNA.
Short Read Libraries (PCR Free)
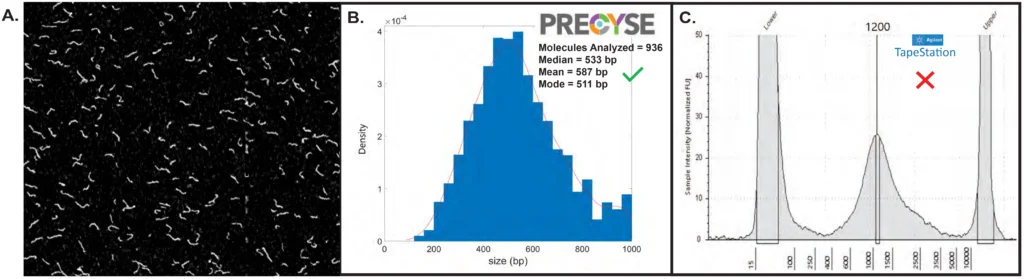
Figure 3. Short Read NGS Library QC
(A,B) (A,B) Example partial frame image from PRECYSE and corresponding sizing analysis of a short read Mouse PCR free Illumina library generated by The Jackson Laboratory. 936 molecules analyzed produced a median of 533 bp, mean of 587 bp and mode at 511 bp.
(C) Agilent TapeStation electropherogram of same PCR Free Illumina library showed a mode of 1200 bp. Slow migration of molecules on the Tapestaion due to Y-adaptors overestimated true sizing. PRECYSE sizing is not impacted by adaptors and secondary structure, and accurately sizes the PCR Free Illumina library.
Long Read Libraries
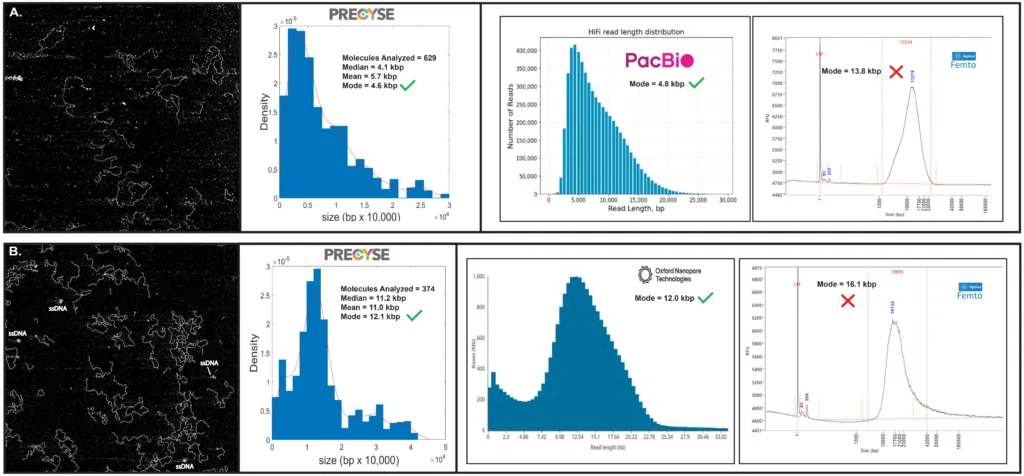
Figure 4. Long Read NGS Library QC
PacBio and Oxford Nanopore (ONT) library QC comparison between PRECYSE and Agilent Femto. All QC and sequencing performed by Harvard Bauer Center.
(A) PacBio final library QC of mouse gDNA prior to polymerase addition generated a mode of 4.6 kbp from 629 molecules analyzed by PRECYSE, Agilent Femto of 13.8 kbp. PacBio read distribution produced a mode of 4.8 kbp, strongly correlating to PRECYSE, while the Femto overestimated sizing by 188%. Additional visual qualitative QC offered only by PRECYSE showed the PacBio library to contain a high concentration of particulates, observed by the white specs.
(B) Oxford Nanopore library QC of Pycnogonida (Sea Spider) gDNA after shearing and cleanup just prior to sequencing. PRECYSE generated a mode of 12.1 kbp, Agilent Femto of 16.1 kbp. Final ONT read length distribution produced a mode of 12.0 kbp. Strong sizing alignment between PRECYSE and ONT, while the Femto overestimated sizing by 34%.
Additional visual qualitative QC offered only by PRECYSE showed the ONT library contained less particulate than the PacBio mouse sample but had some amount of ssDNA.
Conclusion